Compare Protein Synthesis In Prokaryotes And Eukaryotes
Juapaving
Apr 03, 2025 · 6 min read
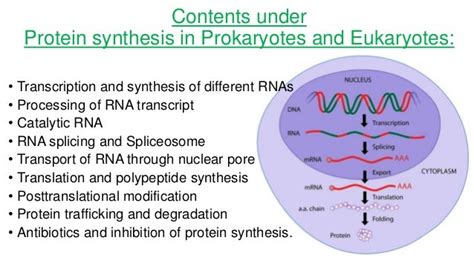
Table of Contents
Comparing Protein Synthesis in Prokaryotes and Eukaryotes: A Detailed Analysis
Protein synthesis, the fundamental process of translating genetic information into functional proteins, is a cornerstone of life. While the basic principles remain consistent across all living organisms, significant differences exist between prokaryotic and eukaryotic protein synthesis. These differences reflect the inherent complexities of eukaryotic cells and have profound implications for various biological processes, including gene regulation and drug targeting. This comprehensive analysis delves into the intricacies of protein synthesis in both prokaryotes and eukaryotes, highlighting key similarities and differences.
Similarities in Prokaryotic and Eukaryotic Protein Synthesis
Despite their structural and organizational differences, prokaryotic and eukaryotic protein synthesis share some fundamental similarities. Both processes involve:
1. Central Dogma: Transcription and Translation
Both prokaryotes and eukaryotes adhere to the central dogma of molecular biology: DNA → RNA → Protein. Genetic information encoded in DNA is first transcribed into messenger RNA (mRNA), which then serves as a template for protein synthesis during translation. This fundamental sequence ensures the faithful transmission of genetic information.
2. Genetic Code Universality
The genetic code, which dictates the correspondence between mRNA codons (three-nucleotide sequences) and amino acids, is largely universal. This means that the same codons specify the same amino acids in both prokaryotes and eukaryotes, with only minor exceptions. This universality underscores the evolutionary conservation of this essential biological process.
3. Ribosomal Machinery
Both prokaryotes and eukaryotes utilize ribosomes, complex molecular machines composed of ribosomal RNA (rRNA) and proteins, to synthesize proteins. Ribosomes facilitate the accurate binding of mRNA and transfer RNA (tRNA), ensuring the correct amino acid sequence is incorporated into the growing polypeptide chain. While the ribosomal structures differ slightly in size and composition between the two domains, their fundamental function remains conserved.
4. tRNA and Aminoacyl-tRNA Synthetase
Transfer RNAs (tRNAs) are essential adaptor molecules that carry specific amino acids to the ribosome based on the mRNA codon sequence. Aminoacyl-tRNA synthetases are enzymes responsible for attaching the correct amino acid to its corresponding tRNA. These processes are remarkably similar in prokaryotes and eukaryotes, ensuring accurate amino acid incorporation during protein synthesis.
Key Differences in Prokaryotic and Eukaryotic Protein Synthesis
Despite the similarities, several crucial differences distinguish prokaryotic and eukaryotic protein synthesis. These differences stem from the structural and organizational disparities between prokaryotic and eukaryotic cells.
1. Transcription and Translation Coupling
In prokaryotes, transcription and translation are coupled processes. Since prokaryotes lack a nucleus, ribosomes can bind to mRNA and initiate translation even before transcription is complete. This allows for a rapid and efficient protein synthesis process. In contrast, eukaryotic transcription and translation are spatially and temporally separated. Transcription occurs in the nucleus, while translation occurs in the cytoplasm. This separation allows for extensive post-transcriptional processing of eukaryotic mRNA, including splicing, capping, and polyadenylation, which are absent in prokaryotes.
2. mRNA Processing
Eukaryotic mRNA undergoes significant post-transcriptional processing before it is exported from the nucleus for translation. This processing includes:
- 5' capping: A 7-methylguanosine cap is added to the 5' end of the mRNA, protecting it from degradation and facilitating ribosome binding.
- Splicing: Introns, non-coding sequences within the mRNA, are removed, and exons, coding sequences, are joined together.
- 3' polyadenylation: A poly(A) tail, a long string of adenine nucleotides, is added to the 3' end of the mRNA, enhancing stability and facilitating translation.
Prokaryotic mRNA, on the other hand, does not undergo these extensive processing steps. It is typically translated directly after transcription.
3. Ribosomal Structure and Function
While both prokaryotes and eukaryotes use ribosomes for protein synthesis, their ribosomal structures differ slightly. Prokaryotic ribosomes (70S) are smaller than eukaryotic ribosomes (80S). This difference in size and composition is exploited by certain antibiotics, which selectively target prokaryotic ribosomes without affecting eukaryotic ribosomes. This selective toxicity forms the basis of many antibacterial drugs.
4. Initiation Factors
The initiation of translation differs significantly between prokaryotes and eukaryotes. Prokaryotes use a single initiation factor (IF3) to bind to the 30S ribosomal subunit, while eukaryotes use several initiation factors (eIFs) that participate in a more complex process of assembling the initiation complex. This complexity allows for more stringent regulation of translation initiation in eukaryotes.
5. Elongation Factors
Elongation, the process of adding amino acids to the growing polypeptide chain, also involves distinct sets of elongation factors in prokaryotes (EF-Tu, EF-Ts, EF-G) and eukaryotes (eEF1α, eEF1βγ, eEF2). These factors play crucial roles in tRNA binding, peptide bond formation, and translocation along the mRNA.
6. Termination Factors
Translation termination is also different. In prokaryotes, release factors RF1, RF2, and RF3 recognize stop codons and mediate the release of the completed polypeptide chain. Eukaryotes, however, use a single release factor, eRF1.
7. Location of Protein Synthesis
In prokaryotes, protein synthesis occurs in the cytoplasm, often simultaneously with transcription. In eukaryotes, transcription occurs in the nucleus, and the mature mRNA is exported to the cytoplasm for translation. This spatial separation allows for additional regulatory checkpoints and contributes to the overall complexity of eukaryotic protein synthesis.
8. Post-translational Modifications
Eukaryotic proteins often undergo extensive post-translational modifications, including glycosylation, phosphorylation, and ubiquitination. These modifications can alter protein function, localization, and stability. While prokaryotic proteins can also be modified, the extent and complexity of post-translational modification are generally less elaborate than in eukaryotes.
9. Regulation of Protein Synthesis
Regulation of protein synthesis is more complex in eukaryotes compared to prokaryotes. Eukaryotes utilize various mechanisms to control gene expression at multiple levels, including transcriptional regulation, mRNA processing, and translational control. Prokaryotic regulation relies more heavily on transcriptional control, particularly through operon systems.
Implications of the Differences
The differences in prokaryotic and eukaryotic protein synthesis have significant implications for various biological processes:
-
Antibiotic development: The differences in ribosomal structure and function between prokaryotes and eukaryotes have been exploited to develop antibiotics that selectively target prokaryotic ribosomes. These antibiotics are effective against bacterial infections without harming the host.
-
Gene expression regulation: The greater complexity of eukaryotic protein synthesis enables finer control of gene expression at multiple levels. This complexity allows for highly regulated and coordinated cellular processes.
-
Disease mechanisms: Dysregulation of protein synthesis is implicated in various diseases, including cancer and neurological disorders. Understanding the differences in protein synthesis between prokaryotes and eukaryotes can help unravel the molecular basis of these diseases.
-
Biotechnology applications: The ability to manipulate protein synthesis pathways in both prokaryotes and eukaryotes has significant applications in biotechnology, including the production of recombinant proteins for therapeutic use.
Conclusion
Protein synthesis is a fundamental process that is remarkably similar yet distinctly different in prokaryotes and eukaryotes. While the basic principles of transcription and translation are conserved, the complexity of eukaryotic cells necessitates additional regulatory mechanisms and post-transcriptional processing steps. Understanding these differences is crucial for various aspects of biology, medicine, and biotechnology. The differences in ribosomal structure, initiation and elongation factors, mRNA processing, and post-translational modifications all contribute to the distinct features of protein synthesis in these two domains of life, making it a rich and fascinating area of ongoing research. Further investigation into these intricacies will undoubtedly lead to breakthroughs in numerous fields, from developing novel therapeutics to understanding the intricacies of life itself.
Latest Posts
Latest Posts
-
What Is The Least Common Denominator Of 12 And 16
Apr 04, 2025
-
How Many Valence Electrons Are In Chlorine
Apr 04, 2025
-
Bar That Is Free To Pivot About A Fixed Point
Apr 04, 2025
-
A Reaction Is Always Spontaneous If
Apr 04, 2025
-
How Many Feet Is 117 Inches
Apr 04, 2025
Related Post
Thank you for visiting our website which covers about Compare Protein Synthesis In Prokaryotes And Eukaryotes . We hope the information provided has been useful to you. Feel free to contact us if you have any questions or need further assistance. See you next time and don't miss to bookmark.
